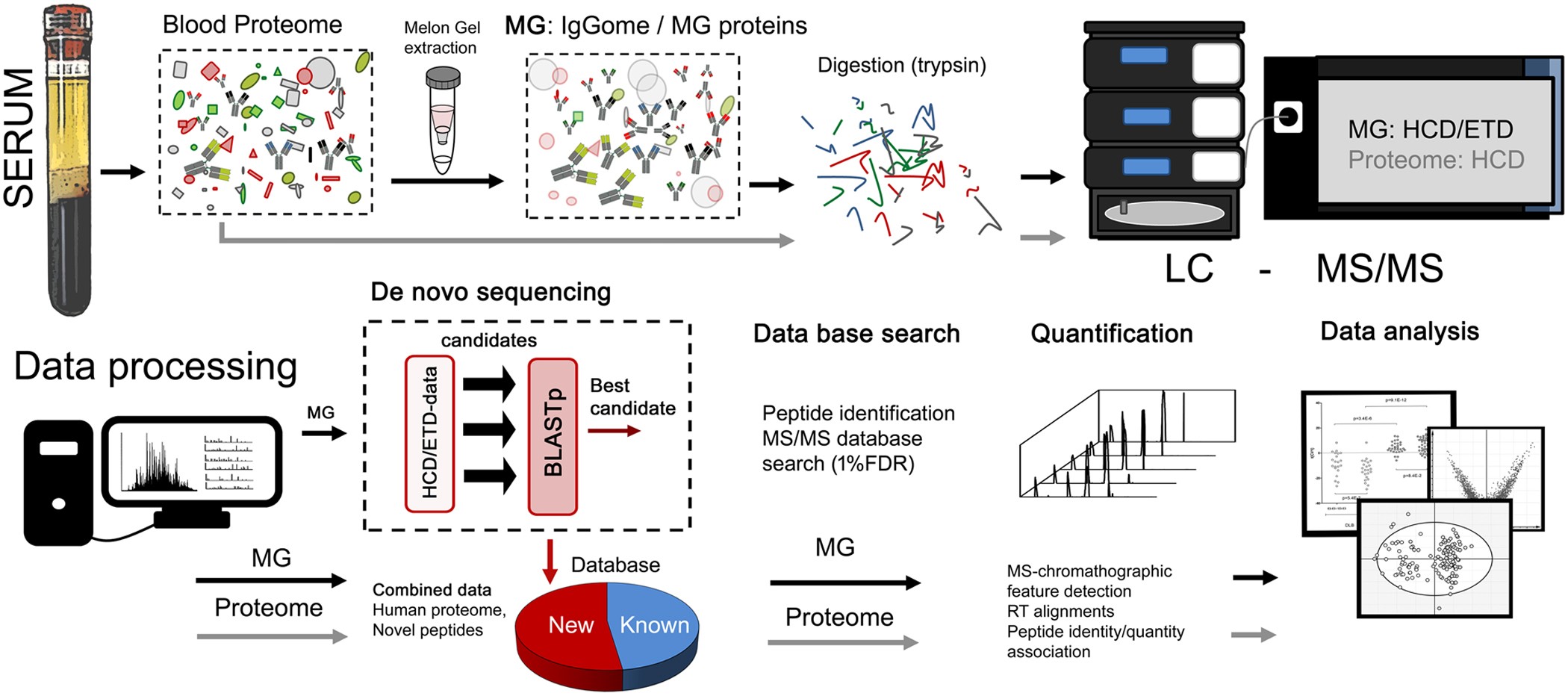
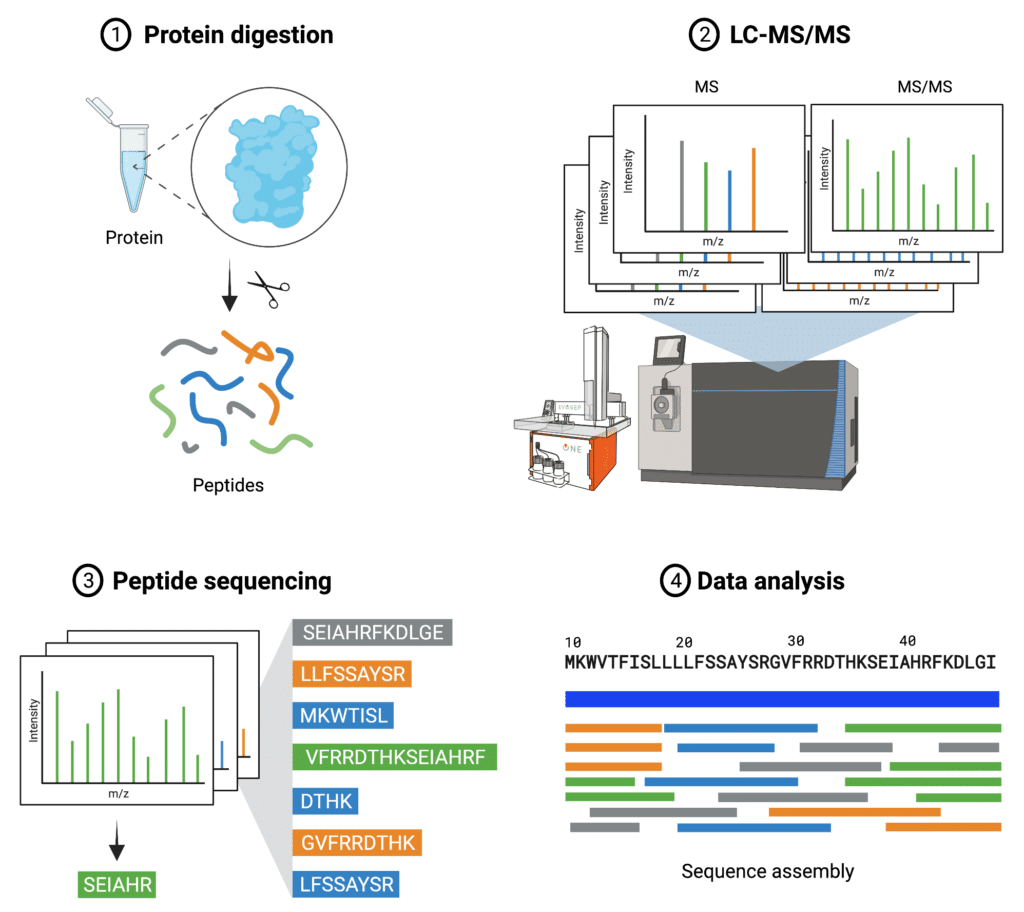
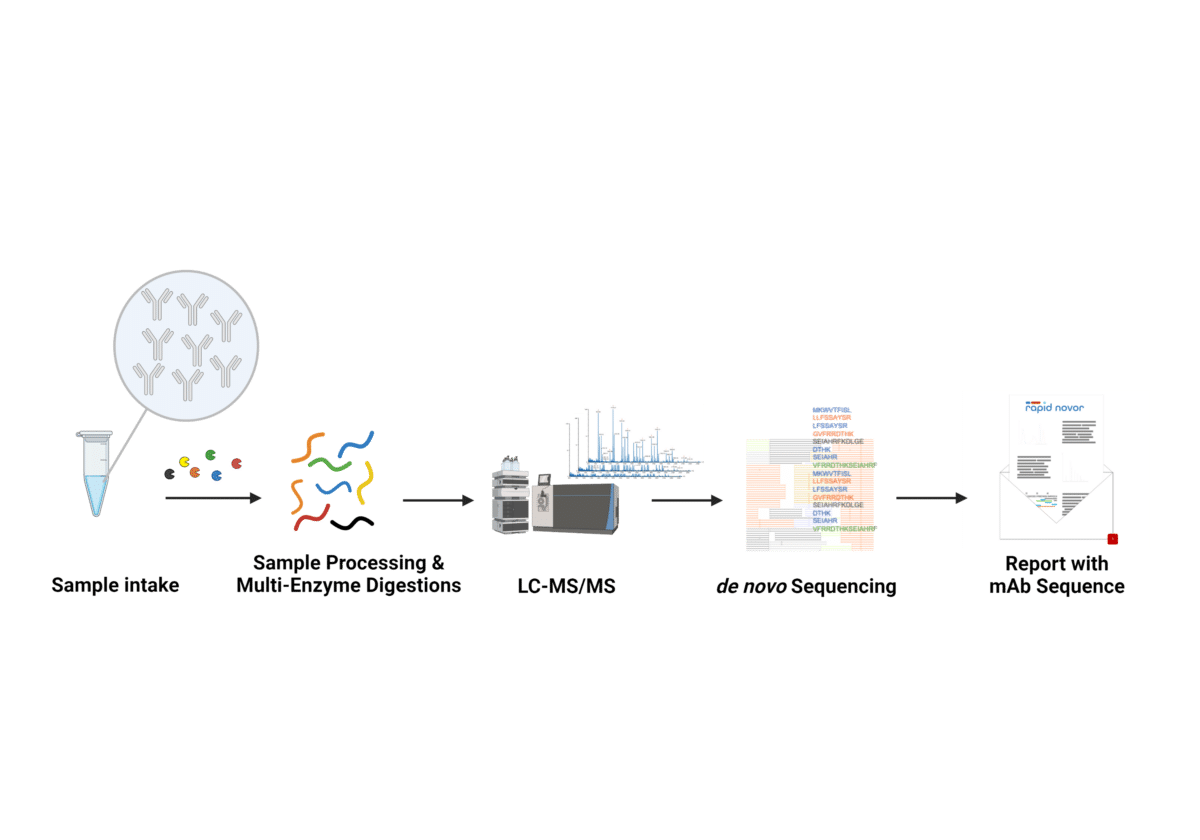
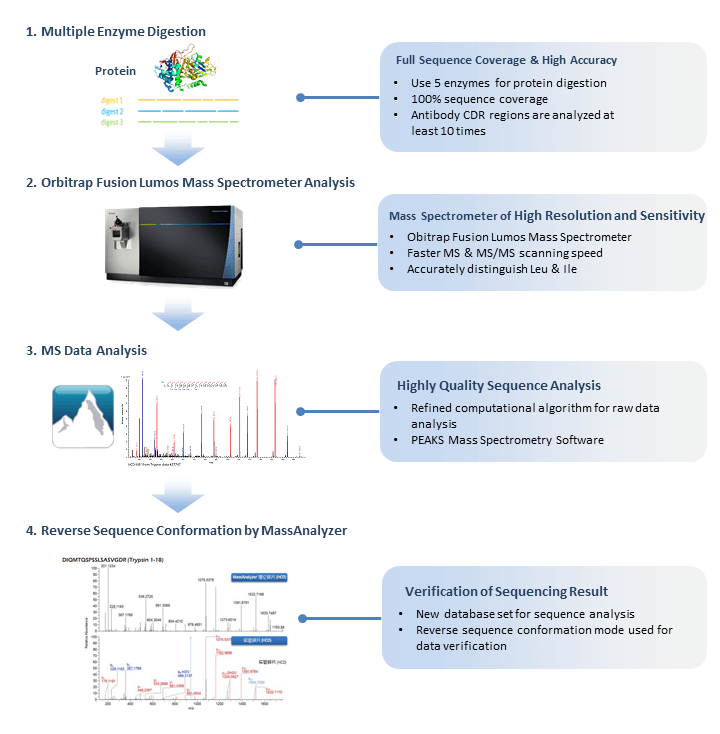
Complementary Methods for de Novo Monoclonal Antibody Sequencing to Achieve Complete Sequence Coverage | Journal of Proteome Research

Precision De Novo Peptide Sequencing Using Mirror Proteases of Ac-LysargiNase and Trypsin for Large-scale Proteomics - ScienceDirect
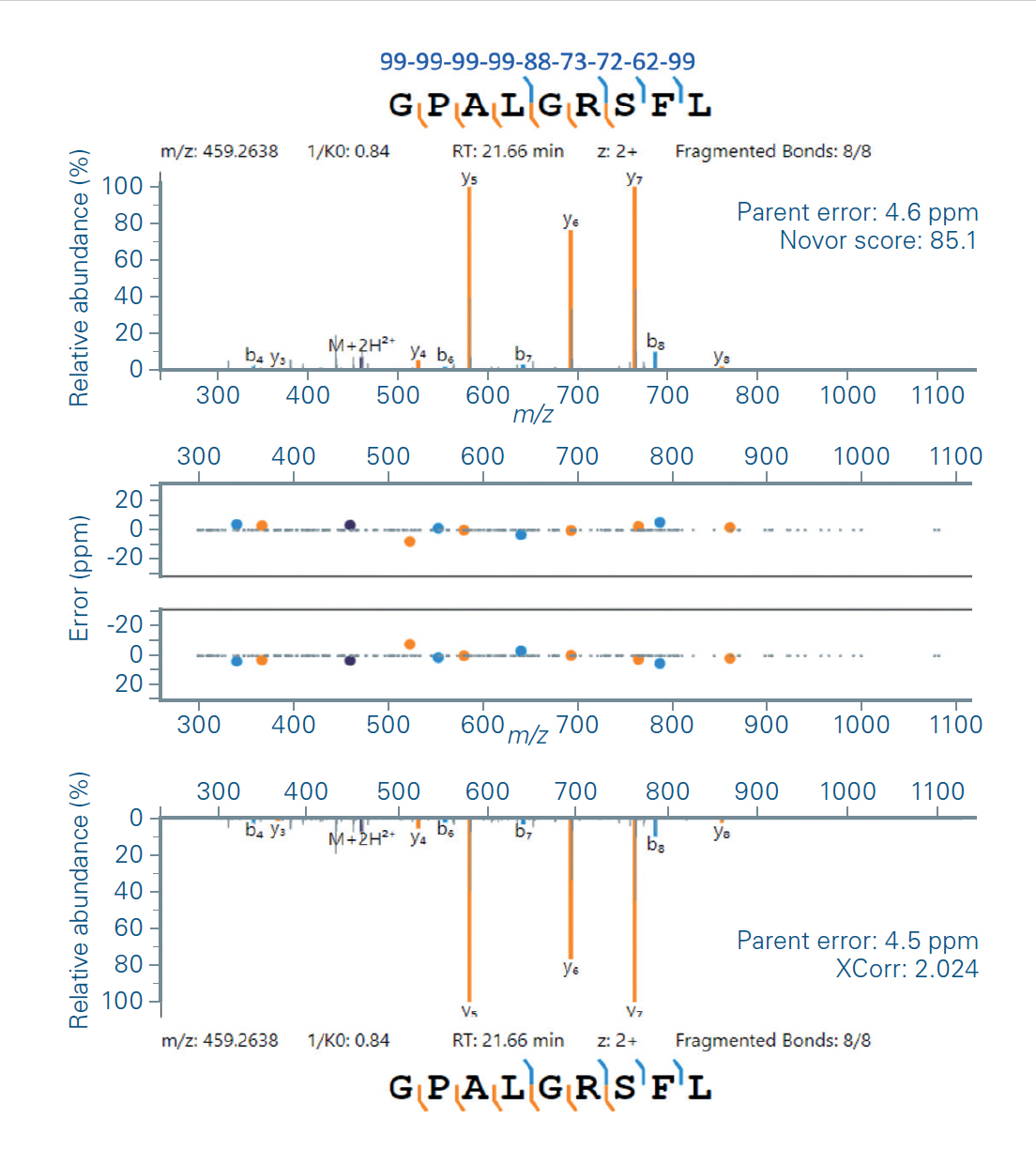
Bruker Launches de novo Sequencing for Immunopeptidomics, Library-Free dia-PASEF, Mass Dynamics Knowledge Visualization | Bruker

A Potential Golden Age to Come—Current Tools, Recent Use Cases, and Future Avenues for De Novo Sequencing in Proteomics - Muth - 2018 - PROTEOMICS - Wiley Online Library
![PDF] Precision De Novo Peptide Sequencing Using Mirror Proteases of Ac-LysargiNase and Trypsin for Large-scale Proteomics* | Semantic Scholar PDF] Precision De Novo Peptide Sequencing Using Mirror Proteases of Ac-LysargiNase and Trypsin for Large-scale Proteomics* | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/f2062e0c44ed5807b4c467e504817018744a86de/6-Figure2-1.png)
PDF] Precision De Novo Peptide Sequencing Using Mirror Proteases of Ac-LysargiNase and Trypsin for Large-scale Proteomics* | Semantic Scholar

Deep learning enables de novo peptide sequencing from data-independent-acquisition mass spectrometry | Nature Methods

Application of de Novo Sequencing to Large-Scale Complex Proteomics Data Sets | Journal of Proteome Research

Deep learning enables de novo peptide sequencing from data-independent-acquisition mass spectrometry | Nature Methods
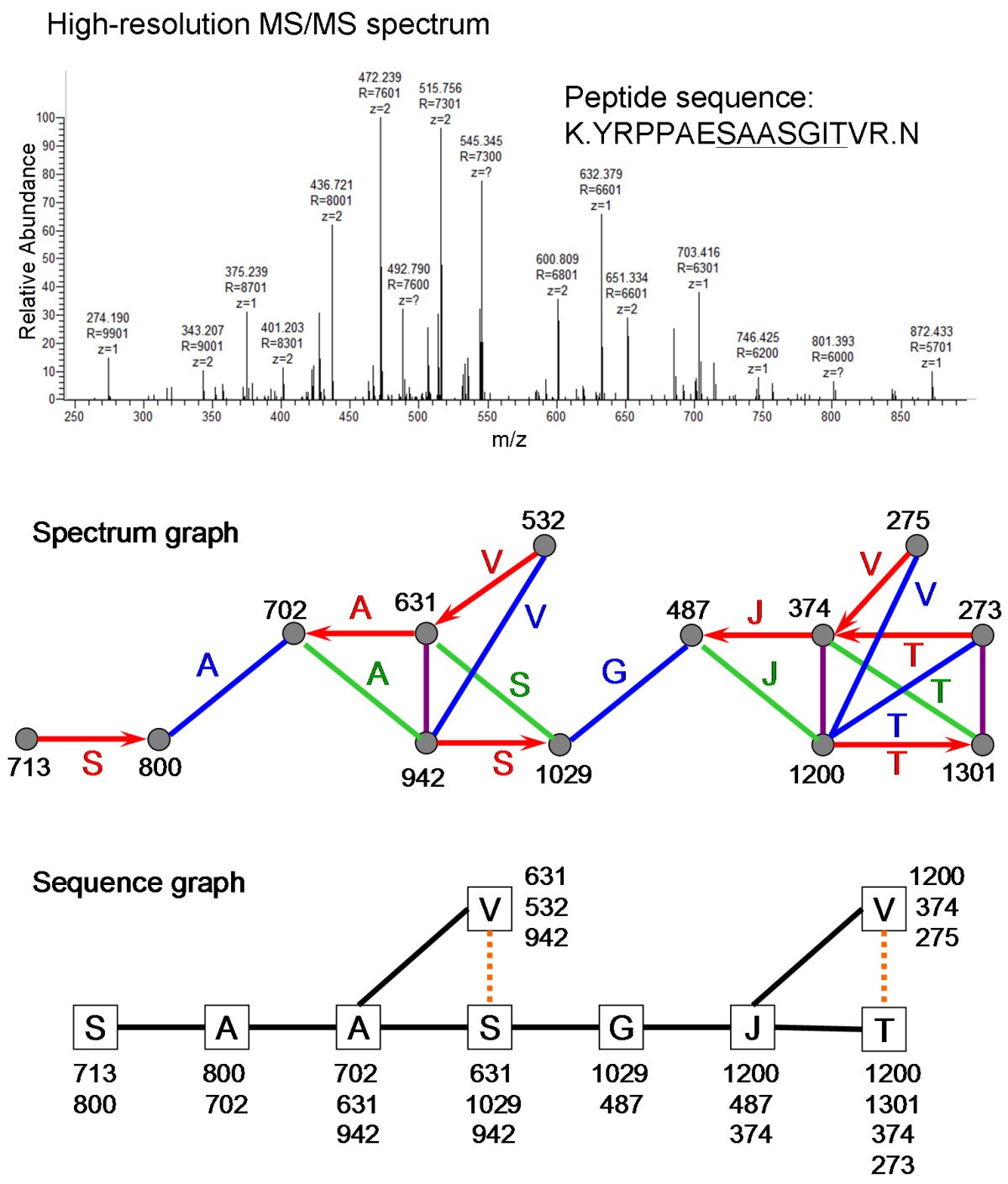
A high-throughput de novo sequencing approach for shotgun proteomics using high-resolution tandem mass spectrometry | BMC Bioinformatics | Full Text