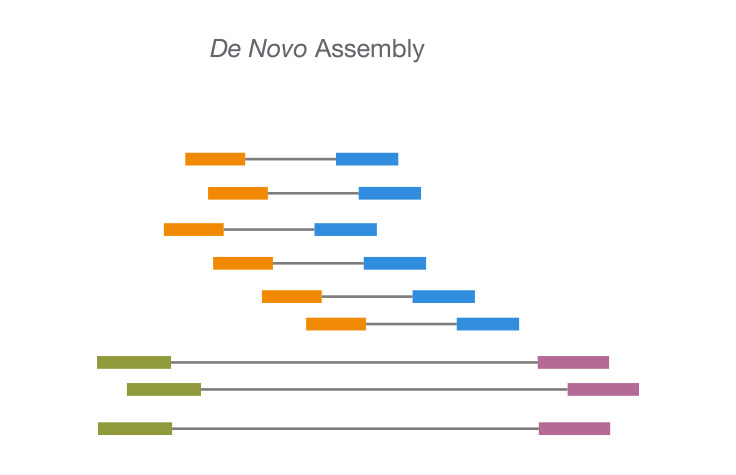
PDF) De novo assembly and functional annotation of blood transcriptome of loggerhead turtle, and in silico characterization of peroxiredoxins and thioredoxins
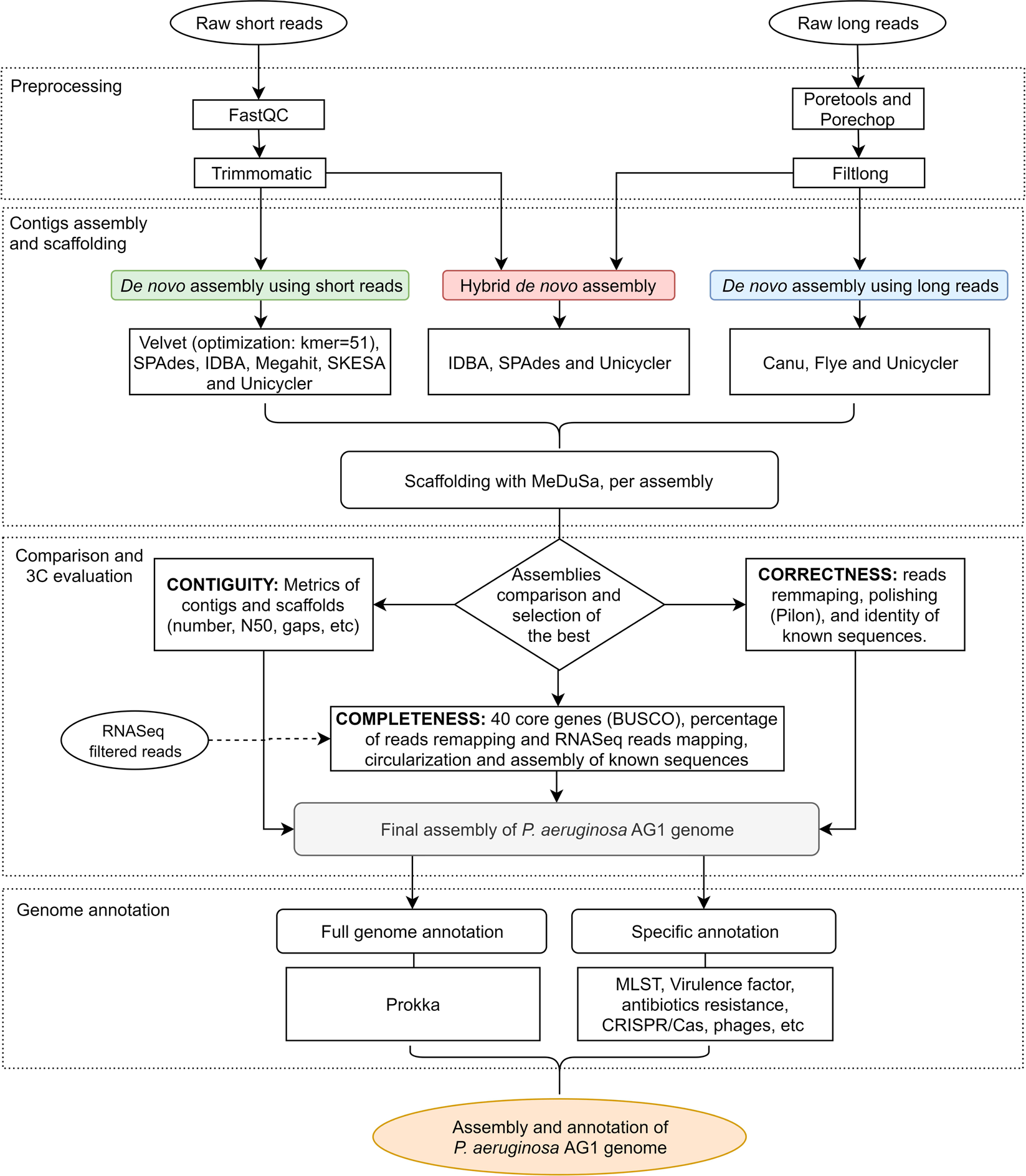
High quality 3C de novo assembly and annotation of a multidrug resistant ST-111 Pseudomonas aeruginosa genome: Benchmark of hybrid and non-hybrid assemblers | Scientific Reports

Mechanism for De Novo RNA Synthesis and Initiating Nucleotide Specificity by T7 RNA Polymerase - ScienceDirect

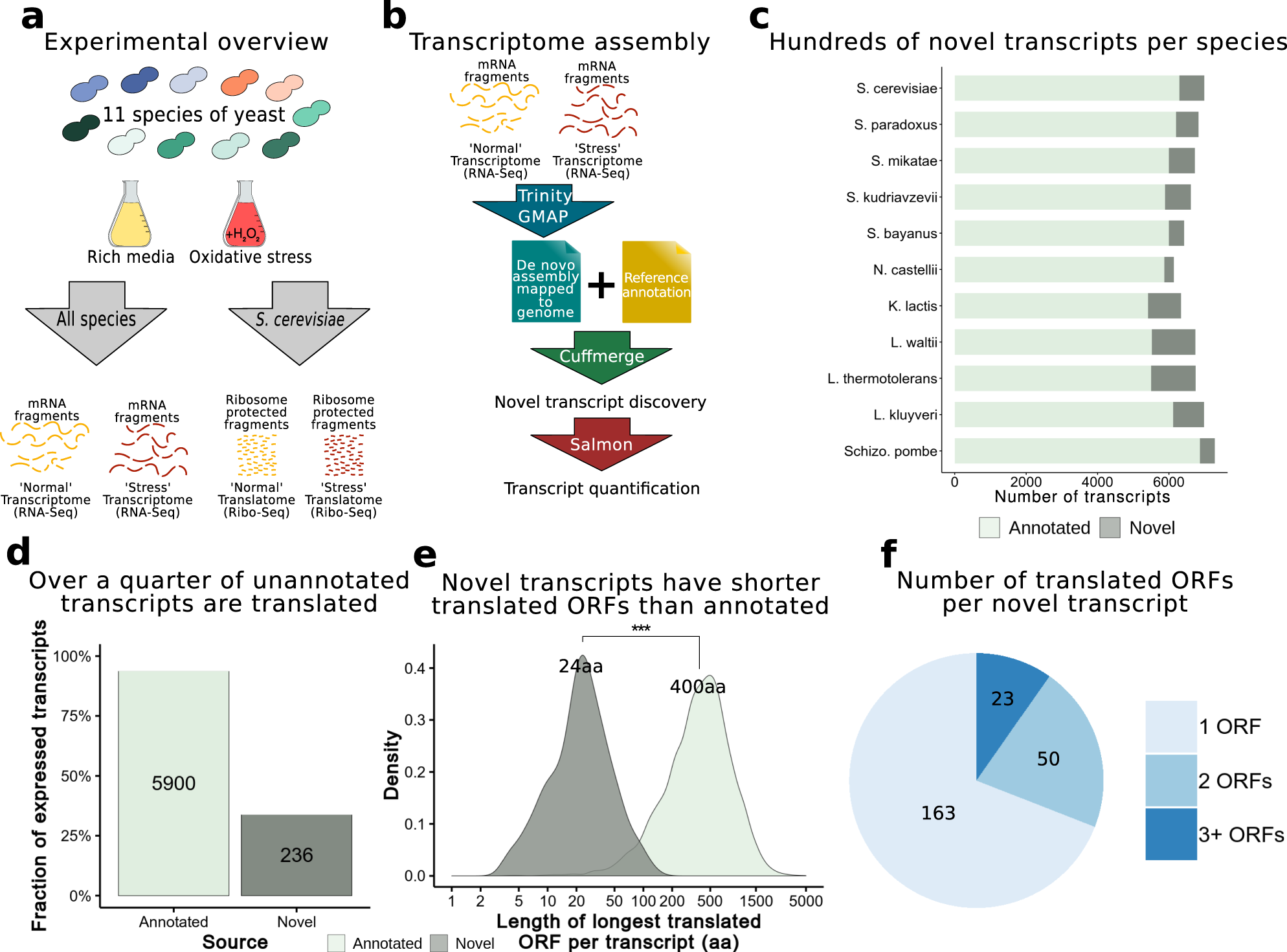
De Novo Transcriptome Sequencing of Ramonda serbica: Identification of the Candidate Genes Involved in the Desiccation Tolerance

Mustafa Kansiz en LinkedIn: Spatiotemporal Heterogeneity of De Novo Lipogenesis in Fixed and Living…
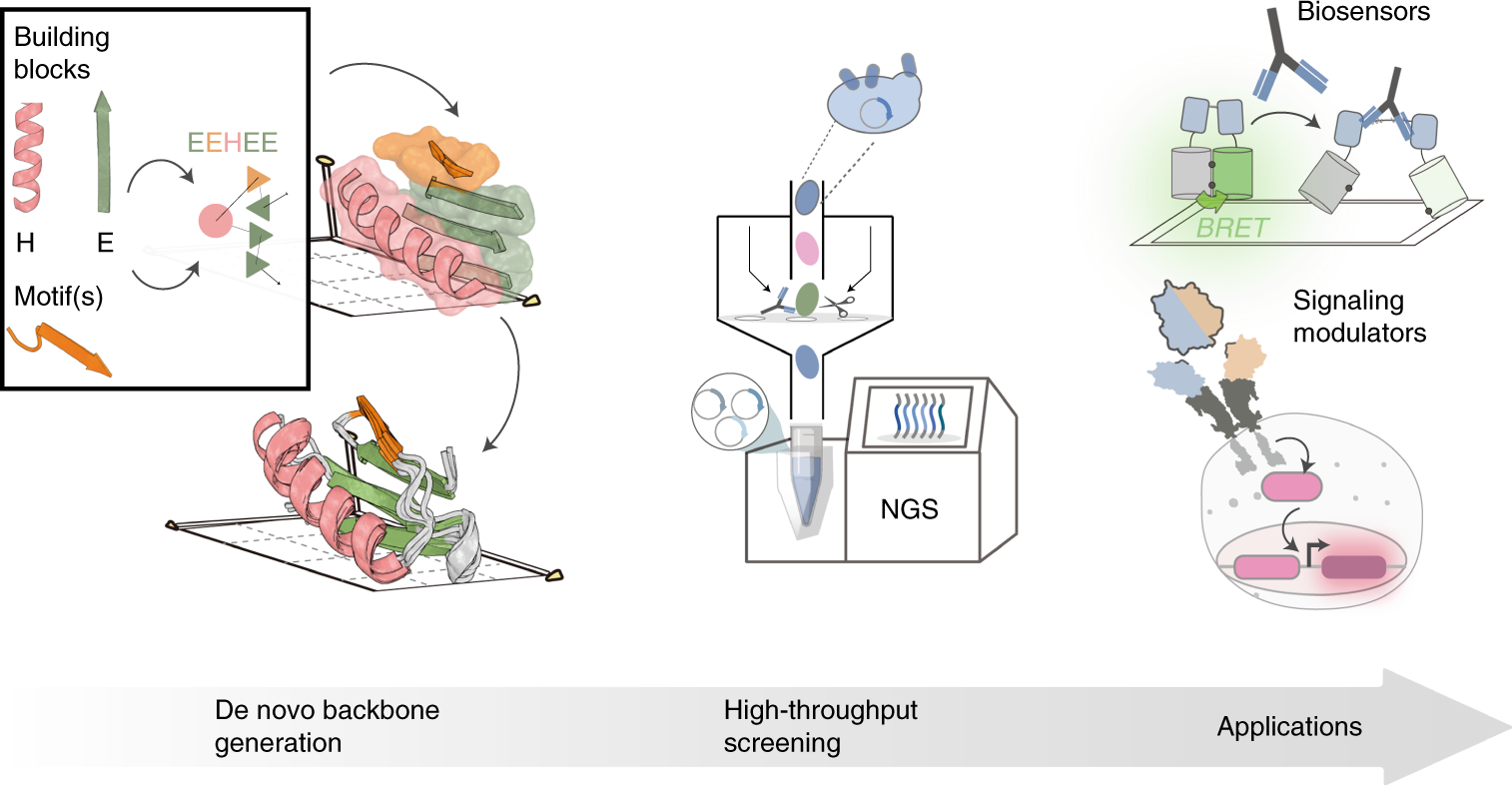
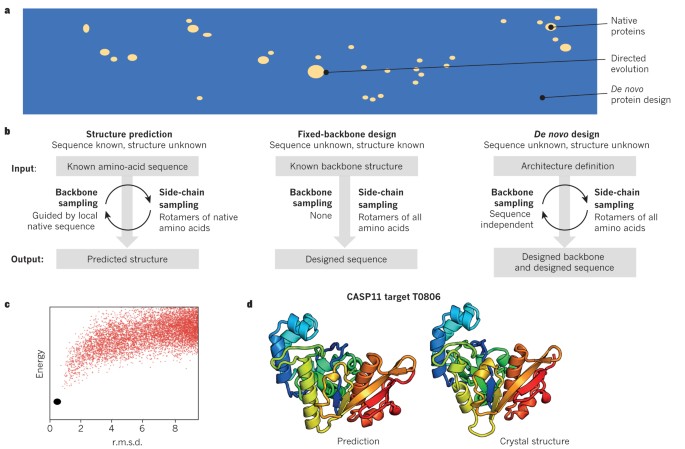
Bottom-up de novo design of functional proteins with complex structural features | Nature Chemical Biology